The HESI COMPrehensive Allergen REsource (COMPARE) Steering Team will meet to plan for the upcoming COMPARE 2024 review cycle.
HESI COMPARE Database Annual Meeting
March 6, 2024 – March 7, 2024
HESI COMPARE Database, Virtual

The PATB is committed to advancing the scientific understanding of the relevant parameters defining allergenic proteins and protein toxins, and encouraging the development of reliable and accurate methodologies for characterizing the allergenic potential of novel proteins in order to leverage the potential of bioinformatics approaches in accomplishing these efforts.
Renaming the Committee: PATC to PATB
Formerly the Protein Allergenicity Technical Committee (PATC), the committee has adopted a new designation, the Protein Allergens, Toxins and Bioinformatics (PATB) Committee, to embrace a broader scope, expertise, interests, and collaborators in the fields of bioinformatics and protein toxins, while strengthening its core focus in allergenicity.
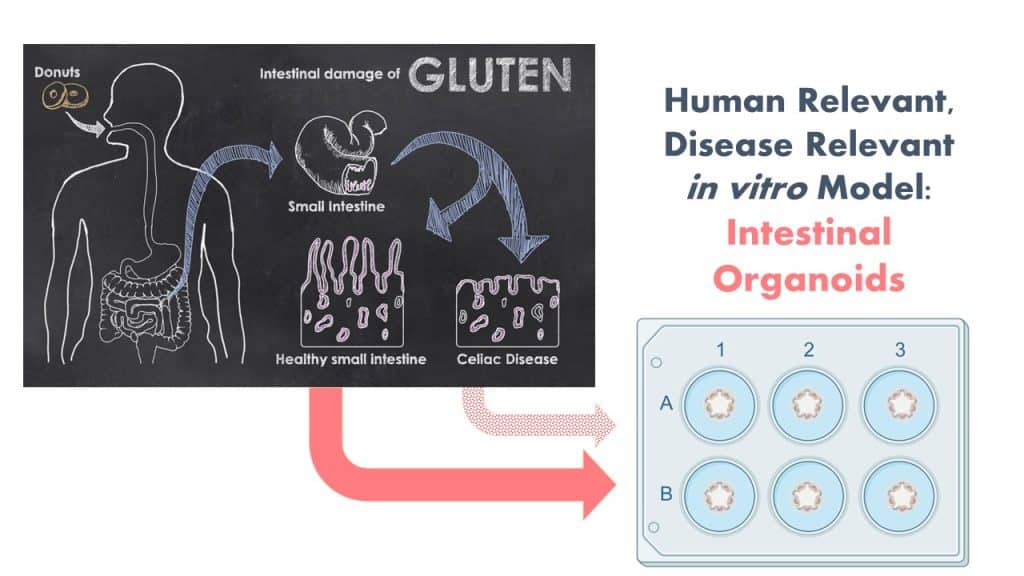
The Protein Allergens, Toxins and Bioinformatics (PATB) Committee is seeking partners to help launch a new project to advance methods for the assessment of gluten-like peptide sequences that could trigger Celiac disease (CD). The proposed pilot study will test the potential of microbial derived peptides to mimic gluten peptides using organoid models.
The project will have relevance for those involved in the evaluation of genetically modified crops, novel foods, or proteins (e.g., for food ingredients) produced via precision fermentation or microbial production systems as well as those with an interest in NAMs for food safety and clinicians/researchers involved in the study of Celiac Disease pathology and immunology.
By involving in vitro NAMs and patient derived materials to generate CD-organoids, this study is anticipated to generate both biological and methodological advancements. We are seeking additional financial support and expertise to ensure that the design and impact of the study is optimized. Your involvement will be enhanced by significant contributions in-kind by a clinical collaborator from Harvard University with expertise in developing these in vitro organoid systems! Peptides will be generated this fall and bench work to launch in early 2024.
As with all HESI projects, the success and impact of this initiative will depend on engaging thought leaders and international stakeholders, from government regulators to clinicians, to crop protection, food biotechnology, and biomedical industry experts, academics, and representatives from other relevant organizations.
Please contact Dr. Lucilia Mouriès at lmouries@hesiglobal.org
Current risk assessment approaches for novel foods include amino acid sequence comparison searches, with comparisons being made to approximately 70 known 9-amino acid celiac epitopes.
However, sequence comparisons have shortcomings. Matching of novel proteins to known celiac epitopes can lead to high false positive rates; for example, many common proteins such as zeins, kafirins, and legumins as well as proteins found in pigs and cows have at least one celiac peptide match.
Furthermore, it has been proposed that changes in the Human gut microbiota can promote CD development in genetically susceptible individuals. A number of microbially derived peptides that share sequence similarities to HLA-DQ2.5-restricted gliadin determinants known to be associated with CD have been identified and shown to activate disease-relevant, gliadin-reactive T-cells isolated from CD patients. Yet, activation of T-cells in the gut is a downstream step in the cascade of events for CD. It is unknown if such peptides, when ingested as foods would pass the intestinal barrier, and reach the T-cells in the gut.
In order to reflect the clinical scenario more wholistically, we propose the use of patient-derived human gut organoids as a model system to determine the celiac potential of a protein containing a single putative CD epitope. Organoids are ideal for this use because they replicate many features of clinical disease in a disease-relevant human in vitro model system.
This research project aims to study the impact of food matrices on the digestibility of proteins and complements the work completed on digestibility in vitro models, by testing whether protocols that take matrices into account would provide a better discrimination of allergens and nonallergens than protocols focusing on purified proteins in solution. In 2020, the experimental work was concluded. The data are being analyzed and a publication is planned in 2021.
The goals of the task force are to (1) investigate approaches for identifying protein toxins and (2) suggest specific guidelines to identify new protein toxins. In 2019, the task force undertook a literature review to identify current approaches being used and leading experts in the topic to design the workshop held in 2020. The workshop covered recent advances in protein toxins biology, described the use of computational biology for protein toxins identification and characterization with in silico approaches, and discussed the applicability of existing tools and resources for safety assessment of novel food biotechnology products.
This working group aims to examine application of an in vitro protocol for identifying specific T-cells and antibodies from nonallergic and allergic patients to pairs of proteins from the same protein family but with different allergenicity. After some delays in the anticipated start date due to the COVID-19 pandemic, the project officially received the green light from the leading institution, Copenhagen University Hospital at Gentofte (Copenhagen, Denmark) in late 2020.
Development and annual update of a database of protein allergen sequences is ongoing. The eighth version of the database was released in January 2024. Throughout the year, the team re-initiates and conducts the 10-month-long process leading to the next annual update of the COMPARE Database. Visit COMPARE at https://comparedatabase.org for more details.




Academic Medical Center, University of Amsterdam

Bayer CropScience
March 6, 2024 – March 7, 2024
HESI COMPARE Database, Virtual
The HESI COMPrehensive Allergen REsource (COMPARE) Steering Team will meet to plan for the upcoming COMPARE 2024 review cycle.
April 30, 2023 – May 4, 2023
St. Louis, MO, USA
Stop by HESI's exhibit booth at ISBR in St. Louis from 30 April - 4 May 2023.
October 21, 2020 – October 22, 2020
Virtual workshop, hosted by the HESI PATB Committee
The HESI Protein Allergens, Toxins, and Bioinformatics (PATB) Committee is hosting a virtual workshop focusing on the state of the science for a wide array of protein toxins classes, MOAs, structures, activity, and general biology. Additionally, the workshop will cover the bioinformatics analysis of protein toxins and how ...
May 22, 2019
Virtual webinar, hosted by the HESI PATB Committee
The Health and Environmental Sciences Institute (HESI) Protein Allergens, Toxins, and Bioinformatics (PATB) Committee is hosting a webinar to discuss emerging opportunities and innovations in the biotechnology and food safety assessment arenas.
October 17, 2018 – October 18, 2018
Copenhagen , Denmark
In the growing field of allergy research, scientists can approach protein characterization and allergy hazard assessment in diverse ways. The Health and Environmental Sciences Institute (HESI) Protein Allergens, Toxins, and Bioinformatics (PATB) Committee supports original basic research that seeks to uncover the ...
October 12, 2016 – October 13, 2016
Rome, Italy
This workshop provided an opportunity to discuss the current state of the science in immunologic mechanisms of non-IgE mediated food allergy with a specific emphasis on use of these data to assess food safety.

Workshop Program Outline (subject to change)
Wednesday, 21 October 2020
Thursday, 22 October 2020
For more information, please contact the PATB Committee's Senior Scientific Program Manager, Dr. Lucilia Mouriès at lmouries@hesiglobal.org.
[post_title] => PATB Committee Protein Toxins Workshop [post_excerpt] => [post_status] => publish [comment_status] => closed [ping_status] => closed [post_password] => [post_name] => patb-committee-protein-toxins-workshop [to_ping] => [pinged] => [post_modified] => 2022-03-07 14:41:34 [post_modified_gmt] => 2022-03-07 14:41:34 [post_content_filtered] => [post_parent] => 0 [guid] => https://hesiglobal.org/?post_type=event&p=22887 [menu_order] => 0 [post_type] => event [post_mime_type] => [comment_count] => 0 [filter] => raw ) [3] => WP_Post Object ( [ID] => 21115 [post_author] => 2 [post_date] => 2019-05-03 09:16:08 [post_date_gmt] => 2019-05-03 14:16:08 [post_content] => REGISTER HERE
REGISTER HERE
The Health and Environmental Sciences Institute (HESI) Protein Allergens, Toxins, and Bioinformatics (PATB) committee will be hosting a webinar to discuss emerging opportunities and innovations in the biotechnology and food safety assessment arenas. See details below!
Details
Speakers:
Advances in agricultural and food biotechnology bring the opportunity to improve the way we grow food and feed crops. As part of the requirements to ensure food safety, it is vital to mitigate potential risks of introducing new allergens or unintended components in the food chain that could induce adverse reactions in humans.
The Health and Environmental Sciences Institute (HESI) Protein Allergens, Toxins, and Bioinformatics (PATB) committee engages international collaborative teams of academics, clinicians, regulators, and industry scientists who conduct research, design and populate public database resources, and convene trainings on the use and evaluation of biotechnology products. The PATB oversees the creation, maintenance, and distribution of the Comprehensive Protein Allergen Resource (COMPARE) Database whose use in conjunction bioinformatic tools, offers an effective means for assessing allergenic potential of novel food proteins.
Webinar attendees will:
 REGISTRATION HERE
MEETING AGENDA
Details
REGISTRATION HERE
MEETING AGENDA
DetailsThere will also be another meeting at the Tivoli Hotel and Congress Center in Copenhagen, Denmark, called FAAM 2018, a EAACI Focused Meeting. FAAM 2018 will take place from 18 -20 October 2018. Please consider attending both events! Find more information on the event here.
About
This educational workshop will cover advances in protein allergy research and inform the use of novel approaches for the identification of potential food allergens. This event will be of interest to government regulators, those involved in food allergy and celiac disease research, and scientists involved in food safety and risk assessment.

Objectives
Evaluate bioinformatics approaches to characterize the allergenicity potential of novel proteins and inform allergy safety assessment.
Discuss real-world examples of: 1) a product development case-study describing the utility of bioinformatics in protein characterization, and 2) a practical application of the EFSA 2017 Guidance.
Provide insights from new research, ranging from modifications of allergen epitopes at the single amino-acid level, to using protein structural modeling for allergenicity prediction, to the relation between enzymes and allergenic activity.
Introduction
Overview of contributions and research of the HESI PATB to Bioinformatics Analysis of the Potential Allergenicity of Novel ProteinsSession I: Bioinformatics Approaches to Protein Characterization and Allergy Safety Assessment
Session II: Other Considerations for Allergen Safety Assessment
The PATB “Allergen Rebuild Project”: can the conservative replacement of amino acids in an allergen epitope affect IgE binding and cross-reactivity?
Session III: Non-IgE Mediated Allergy
Overview of known non-IgE mediated diseases and progress in immunology of non-IgE diseasesRegulatory Toxicology and Pharmacology, 2022
The HESI Protein Allergens, Toxins, and Bioinformatics Committee, and the Society of Toxicology Food Safety Specialty Section co-hosted a virtual workshop titled “From Protein Toxins to Applied Toxicological Testing”. Key outcomes of the workshop are highlighted here.
Frontiers in Allergy, 2022
Our aim was to evaluate the impact of experimental and endogenous food matrices on protease susceptibility of homologous protein pairs with different degrees of allergenicity.
Frontiers in Allergy, 2021
The availability of databases identifying allergenic proteins via a transparent and consensus-based scientific approach is of prime importance to support the safety review of genetically-modified foods and feeds, and public safety in general. Over recent years, screening for potential new allergens sequences has become ...
Clinical and Translational Allergy, 2018
Susceptibility to pepsin digestion of candidate transgene products is regarded an important parameter in the weight-of-evidence approach for allergenicity risk assessment of genetically modified crops. It has been argued that protocols used for this assessment should better reflect physiological ...
Frontiers in Plant Science, 2018
Soybean (Glycine max) is an important food stock, and also considered an allergenic food with at least eight well characterized allergens. However, it is a less prevalent allergen source than many other foods and is rarely life-threatening. Soybean is incorporated into commonly consumed foods, and therefore, the allergens pose ...
Scientific Reports, 2017
Proteins are fundamental to life and exhibit a wide diversity of activities, some of which are toxic. Therefore, assessing whether a specific protein is safe for consumption in foods and feeds is critical. Simple BLAST searches may reveal homology to a known toxin, when in fact the protein may pose no real danger.
hesi@hesiglobal.org
Phone: +1-202-659-8404
Fax: +1-202-659-3859
740 15th Street NW, Suite 600
Washington, DC 20005
Sign up for our monthly e-newsletter.